Create a GAMLSS Distribution
GAMLSS.RdA single class and corresponding methods encompassing all distributions from the gamlss.dist package using the workflow from the distributions3 package.
GAMLSS(family, mu, sigma, tau, nu)Arguments
- family
character. Name of a GAMLSS family provided by gamlss.dist, e.g.,
NOorBIfor the normal or binomial distribution, respectively.- mu
numeric. GAMLSS
muparameter. Can be a scalar or a vector or missing if not part of thefamily.- sigma
numeric. GAMLSS
sigmaparameter. Can be a scalar or a vector or missing if not part of thefamily.- tau
numeric. GAMLSS
tauparameter. Can be a scalar or a vector or missing if not part of thefamily.- nu
numeric. GAMLSS
nuparameter. Can be a scalar or a vector or missing if not part of thefamily.
Value
A GAMLSS distribution object.
Details
The constructor function GAMLSS sets up a distribution
object, representing a distribution from the GAMLSS (generalized additive
model of location, scale, and shape) framework by the corresponding parameters
plus a family attribute, e.g., NO for the
normal distribution or BI for the binomial
distribution. There can be up to four parameters, called mu (often some
sort of location parameter), sigma (often some sort of scale parameter),
tau and nu (other parameters, e.g., capturing shape, etc.).
All parameters can also be vectors, so that it is possible to define a vector of GAMLSS distributions from the same family with potentially different parameters. All parameters need to have the same length or must be scalars (i.e., of length 1) which are then recycled to the length of the other parameters.
Note that not all distributions use all four parameters, i.e., some use just a
subset. In that case, the corresponding arguments in GAMLSS should be
unspecified, NULL, or NA.
For the GAMLSS distribution objects there is a wide range
of standard methods available to the generics provided in the distributions3
package: pdf and log_pdf
for the (log-)density (PDF), cdf for the probability
from the cumulative distribution function (CDF), quantile for quantiles,
random for simulating random variables,
and support for the support interval
(minimum and maximum). Internally, these methods rely on the usual d/p/q/r
functions provided in gamlss.dist, see the manual pages of the individual
families. The methods is_discrete and
is_continuous can be used to query whether the
distributions are discrete on the entire support or continuous on the entire
support, respectively.
Additionally, for some families there is also a crps
method for computing the continuous ranked probability score (CRPS) via the
scoringRules package. This is only available for those families which are
supported by both packages.
See the examples below for an illustration of the workflow for the class and methods.
See also
Examples
if(!requireNamespace("gamlss.dist")) {
if(interactive() || is.na(Sys.getenv("_R_CHECK_PACKAGE_NAME_", NA))) {
stop("not all packages required for the example are installed")
} else q() }
#> Loading required namespace: gamlss.dist
#> Registered S3 methods overwritten by 'gamlss.dist':
#> method from
#> format.GAMLSS topmodels
#> print.GAMLSS topmodels
#> mean.GAMLSS topmodels
#> quantile.GAMLSS topmodels
## package and random seed
library("distributions3")
set.seed(6020)
## three Weibull distributions
X <- GAMLSS("WEI", mu = c(1, 1, 2), sigma = c(1, 2, 2))
X
#> [1] "GAMLSS WEI distribution (mu = 1, sigma = 1)"
#> [2] "GAMLSS WEI distribution (mu = 1, sigma = 2)"
#> [3] "GAMLSS WEI distribution (mu = 2, sigma = 2)"
## moments
mean(X)
#> [1] 1.0000000 0.8862269 1.7724539
variance(X)
#> [1] 1.0000000 0.2146018 0.8584073
## support interval (minimum and maximum)
support(X)
#> min max
#> [1,] 0 Inf
#> [2,] 0 Inf
#> [3,] 0 Inf
is_discrete(X)
#> [1] FALSE FALSE FALSE
is_continuous(X)
#> [1] TRUE TRUE TRUE
## simulate random variables
random(X, 5)
#> r_1 r_2 r_3 r_4 r_5
#> [1,] 1.004812 0.3993868 1.3084904 0.741085 3.209012
#> [2,] 1.023982 0.5593371 0.5440658 1.046042 2.244104
#> [3,] 1.254624 2.8848168 3.0103388 2.166692 2.572731
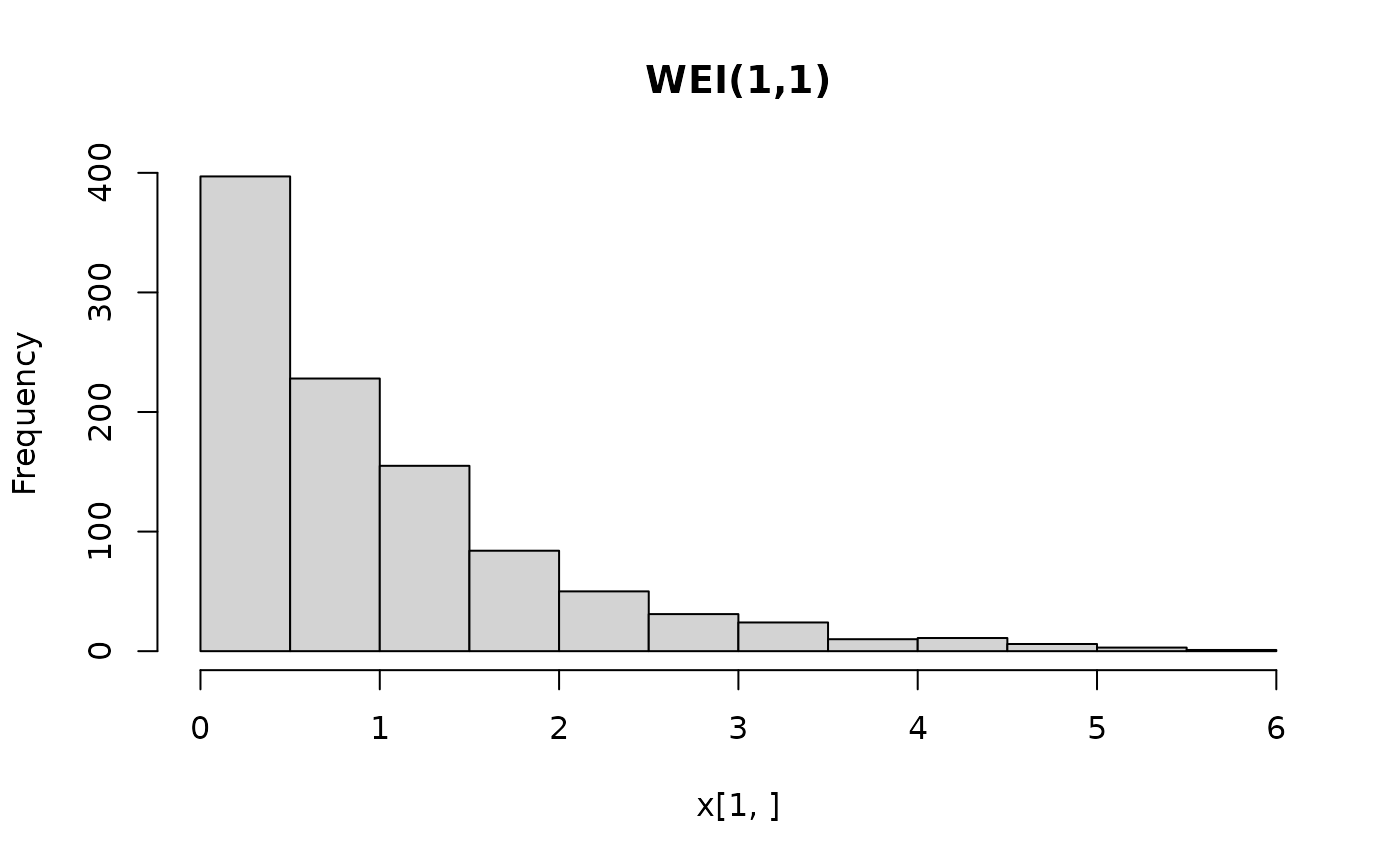
## histograms of 1,000 simulated observations
x <- random(X, 1000)
hist(x[1, ], main = "WEI(1,1)")
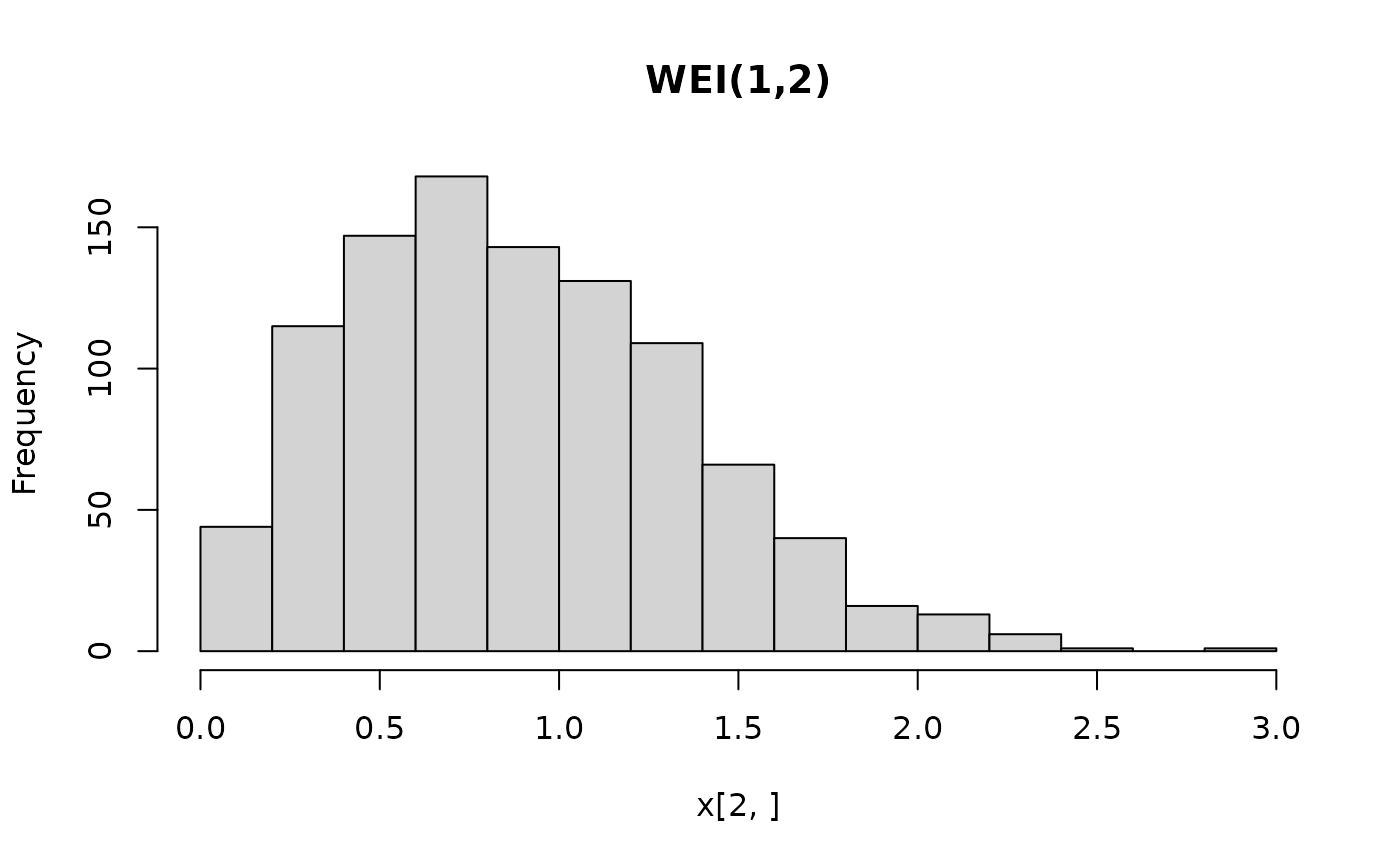
 hist(x[2, ], main = "WEI(1,2)")
hist(x[2, ], main = "WEI(1,2)")
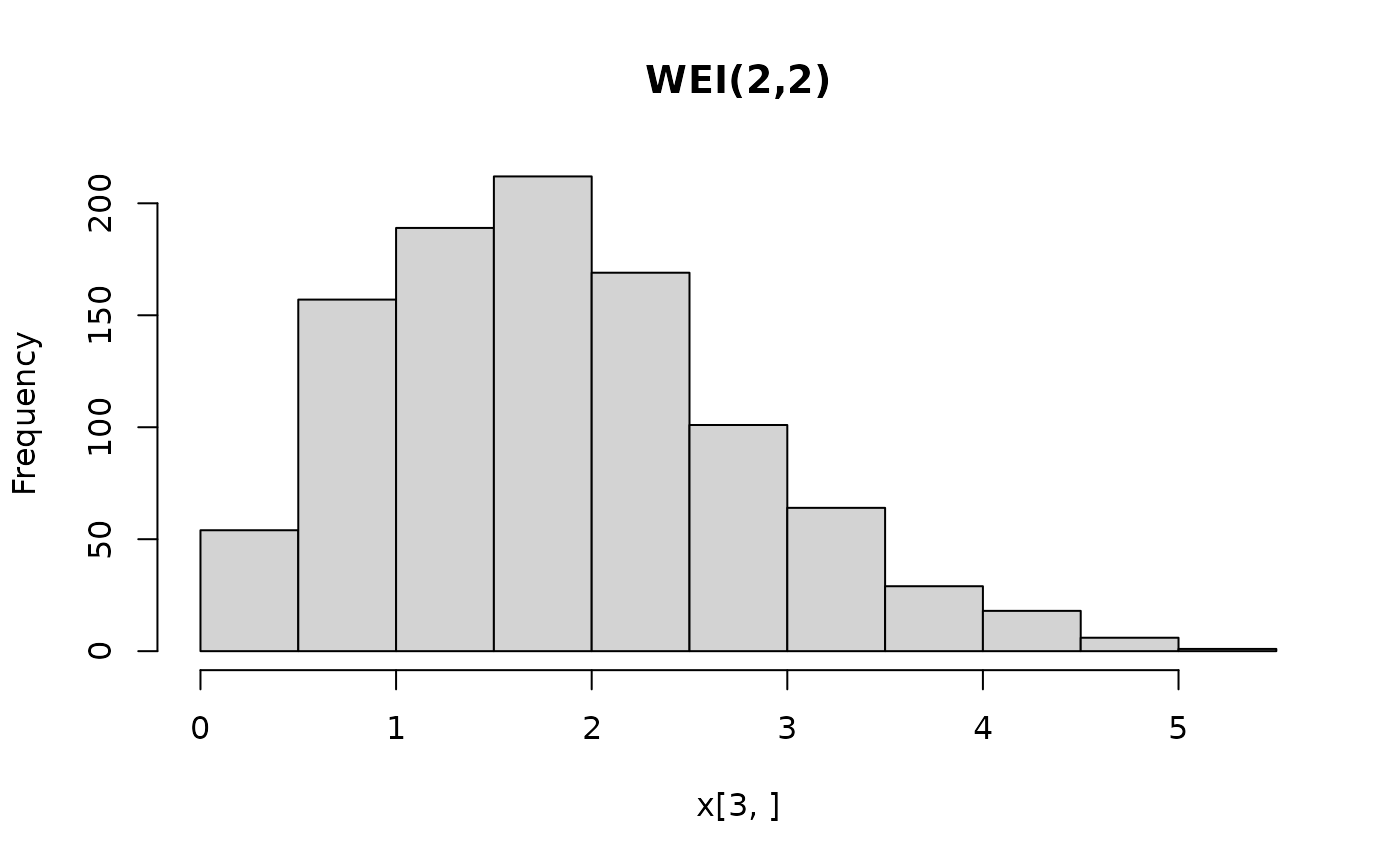
 hist(x[3, ], main = "WEI(2,2)")
hist(x[3, ], main = "WEI(2,2)")
 ## probability density function (PDF) and log-density (or log-likelihood)
x <- c(2, 2, 1)
pdf(X, x)
#> [1] 0.13533528 0.07326256 0.38940039
pdf(X, x, log = TRUE)
#> [1] -2.0000000 -2.6137056 -0.9431472
log_pdf(X, x)
#> [1] -2.0000000 -2.6137056 -0.9431472
## cumulative distribution function (CDF)
cdf(X, x)
#> [1] 0.8646647 0.9816844 0.2211992
## quantiles
quantile(X, 0.5)
#> [1] 0.6931472 0.8325546 1.6651092
## cdf() and quantile() are inverses
cdf(X, quantile(X, 0.5))
#> [1] 0.5 0.5 0.5
quantile(X, cdf(X, 1))
#> [1] 1 1 1
## all methods above can either be applied elementwise or for
## all combinations of X and x, if length(X) = length(x),
## also the result can be assured to be a matrix via drop = FALSE
p <- c(0.05, 0.5, 0.95)
quantile(X, p, elementwise = FALSE)
#> q_0.05 q_0.5 q_0.95
#> [1,] 0.05129329 0.6931472 2.995732
#> [2,] 0.22648023 0.8325546 1.730818
#> [3,] 0.45296046 1.6651092 3.461637
quantile(X, p, elementwise = TRUE)
#> [1] 0.05129329 0.83255461 3.46163677
quantile(X, p, elementwise = TRUE, drop = FALSE)
#> quantile
#> [1,] 0.05129329
#> [2,] 0.83255461
#> [3,] 3.46163677
## compare theoretical and empirical mean from 1,000 simulated observations
cbind(
"theoretical" = mean(X),
"empirical" = rowMeans(random(X, 1000))
)
#> theoretical empirical
#> [1,] 1.0000000 1.0075953
#> [2,] 0.8862269 0.9057806
#> [3,] 1.7724539 1.7587308
## probability density function (PDF) and log-density (or log-likelihood)
x <- c(2, 2, 1)
pdf(X, x)
#> [1] 0.13533528 0.07326256 0.38940039
pdf(X, x, log = TRUE)
#> [1] -2.0000000 -2.6137056 -0.9431472
log_pdf(X, x)
#> [1] -2.0000000 -2.6137056 -0.9431472
## cumulative distribution function (CDF)
cdf(X, x)
#> [1] 0.8646647 0.9816844 0.2211992
## quantiles
quantile(X, 0.5)
#> [1] 0.6931472 0.8325546 1.6651092
## cdf() and quantile() are inverses
cdf(X, quantile(X, 0.5))
#> [1] 0.5 0.5 0.5
quantile(X, cdf(X, 1))
#> [1] 1 1 1
## all methods above can either be applied elementwise or for
## all combinations of X and x, if length(X) = length(x),
## also the result can be assured to be a matrix via drop = FALSE
p <- c(0.05, 0.5, 0.95)
quantile(X, p, elementwise = FALSE)
#> q_0.05 q_0.5 q_0.95
#> [1,] 0.05129329 0.6931472 2.995732
#> [2,] 0.22648023 0.8325546 1.730818
#> [3,] 0.45296046 1.6651092 3.461637
quantile(X, p, elementwise = TRUE)
#> [1] 0.05129329 0.83255461 3.46163677
quantile(X, p, elementwise = TRUE, drop = FALSE)
#> quantile
#> [1,] 0.05129329
#> [2,] 0.83255461
#> [3,] 3.46163677
## compare theoretical and empirical mean from 1,000 simulated observations
cbind(
"theoretical" = mean(X),
"empirical" = rowMeans(random(X, 1000))
)
#> theoretical empirical
#> [1,] 1.0000000 1.0075953
#> [2,] 0.8862269 0.9057806
#> [3,] 1.7724539 1.7587308